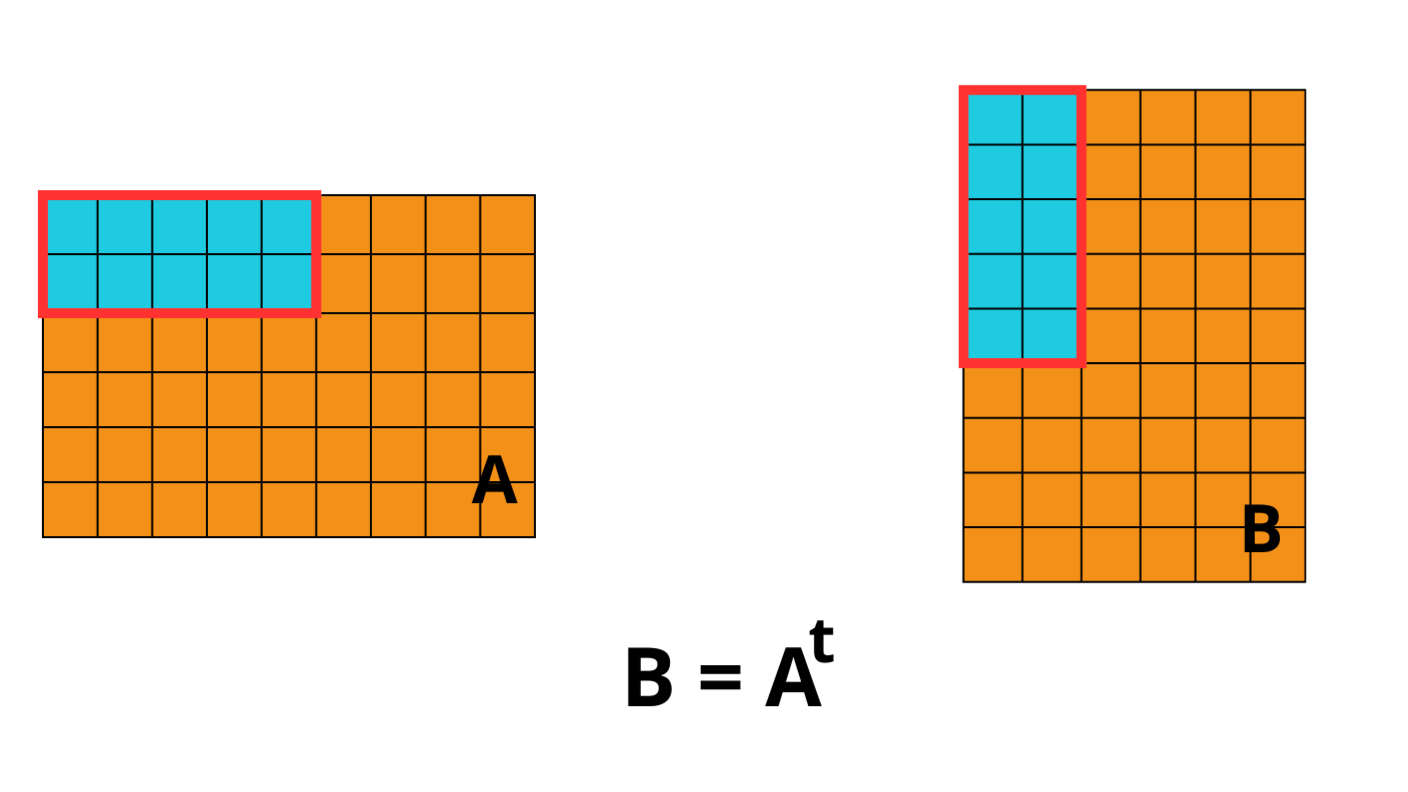
Introducing CTable: a Blosc2-based columnar table
Working with large structured datasets in Python often means choosing between speed and simplicity. The new CTable object has born out of the need for a columnar store that compresses data on the fly, stays close to NumPy, and does not require an external database engine. It is the logical extension of current compressed data storage and computation in Python-Blosc2, brought to tabular datasets.
As compression is paramount in Blosc2 ecosystem, we have chosen a columnar approach because, by placing together data that is similar (values in the same column), it allows for best compression ratios. Column storage also allows for better data management for some cases, like adding, deleting, accessing or replacing entire columns; admittedly, it it also has its own drawbacks, like more costly access along the row axis. Nevertheless, columnar storage is quite common in modern libraries.
Another important piece for CTable is to leverage the extremely efficient compute engine that can operate on compressed data without dropping too much performance (and in some cases, even improving it). That puts the basis for allowing great analytics machinery on top of the CTable object, without the need to decompress entire columns (just small excerpts of them, fitting in CPU caches, is enough).
Last but not least, the CTable object inherits the independence of media storage of underlying structures (NDArray, ObjectArray, ListArray...) so that data can be stored and used straight from memory, disk or the network (coming soon). That means that you can open a data file containing a big CTable and immediately start doing analytics with it without the need to load/parse everything in-memory. Of course, for maximum speed, you may also load everything in-memory too; but as the format is the same, loading/saving is just a matter of copying data from one media to another, without the need for parsing or conversion.
Keep reading to learn more about CTable, its features and how to use it in your projects.
How it works
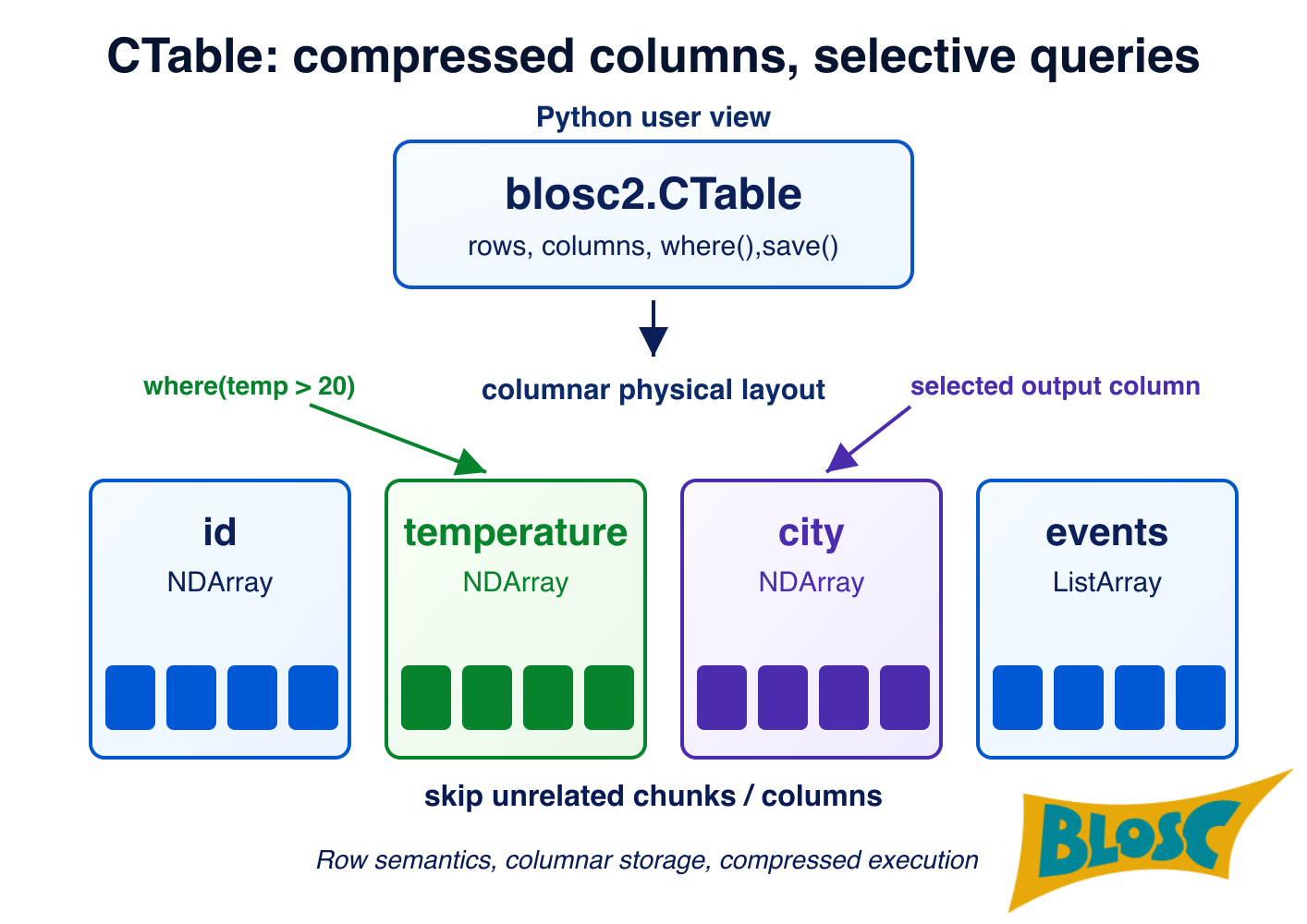
CTable stores each column as an independent blosc2.NDArray, compressed in chunks. Column types are defined through a schema — a plain Python dataclass where each field is annotated with a Blosc2 type spec such as b2.int64(), b2.float32(ge=0), or b2.string(). Specs can carry constraints (e.g. ge=0 for non-negative values) and are compiled into a schema that validates every row on insert, either one at a time via Pydantic or in bulk via vectorized NumPy checks.
Rows are tracked with a boolean tombstone mask: deleting a row simply flips its entry in the mask to False, with no data movement at all. The actual space is reclaimed lazily when you call compact(). Appending is also efficient because the underlying arrays are pre-allocated up front — they only grow when the pre-allocated capacity is exhausted, so there is no resize on every single insert.
Because the data lives in Blosc2 chunks, many queries can skip full chunks entirely. When a chunk's stored metadata (min/max) rules out any match, it is never decompressed. This is where a lot of the query speed comes from, and it is also why explicit indexes (described in Features below) can push performance even further.
Since blosc2.NDArray stores fixed-width binary data, null has no natural representation for integers, floats, or booleans. CTable solves this by letting you declare a column as nullable, and a sentinel value as the null marker is chosen automatically, although you can always use your own. For example, if you are storing ages and sometimes the value is unknown, you can set -1 as the null value since ages are never negative. Aggregates such as .mean() or .std() skip those rows automatically, and helper methods like .is_null() and .null_count() make it easy to work with them.
Not all data fits neatly into fixed-width columns though. Think of a column storing the tags of an article, the purchase history of a user, or the list of measurements taken at a sensor in a given day — each row may have a different number of items. For these cases CTable supports list columns, declared as blosc2.list(item_spec) (e.g. blosc2.list(blosc2.float32())), structured objects via blosc2.struct(item_spec) or completely general objects via b2.object(). These columns are backed by a different storage class internally, one that keeps a compressed stream of items alongside an offsets array to know where each row starts and ends. From the user's perspective they behave like any other column, but each cell holds a Python list instead of a scalar, and individual lists can also be None. Internally, the underlying C-Blosc2 has been improved (and released as 3.0.0) to allow variable-length data in super-chunks in a very efficient (and backward-compatible) way.
Main features
Creation: A CTable can be created in several ways. The most direct is declaring a typed schema as a dataclass and passing it to the constructor. You can also build a CTable from existing data —
from_arrow()andfrom_csv()import Arrow tables and CSV files respectively, inferring or mapping types automatically. Finally,copy()produces a new independent CTable from an existing one, already compacted.Modification: Appending a single row uses
append(), while bulk insertion usesextend(). Deleting rows sets their mask entry toFalseand is essentially free. Columns can be added with a default value or dropped and renamed at any time. Beyond stored columns, CTable also supports two kinds of virtual columns: computed columns are evaluated on-the-fly from an expression over other columns and never touch storage; materialized columns look like stored columns but are auto-filled automatically during everyextend().Querying:
where(expr)filters rows and returns a view — a new CTable object that shares the same column arrays as the parent but carries its own mask. No data is copied; only the mask is computed. Views block structural changes (adding/dropping columns, deleting rows) but do allow writing values to existing cells.select(cols)gives a column-projection view in the same spirit. Both can be made into a fully independent mutable table withcopy(). Aggregates (sum(),mean(),std(),min(),max(), ...) andsort_by()also work on views and respect the mask.Indexing: For workloads that repeatedly query the same column, CTable supports three index flavors:
FULL(sorted positions array, best for range and comparison queries),BUCKET(hash-based, best for equality lookups), andPARTIAL(a lighter-weight sorted structure). Once an index is created,where()uses it automatically when the query can benefit from it. Indexes are persisted alongside the table and survive.b2zround-trips.Persistence: Tables can live fully in memory or be backed by files on disk.
save()writes an in-memory table to a directory or.b2zarchive.CTable.open()attaches directly to an on-disk table for reading or writing without loading everything into RAM.CTable.load()copies the on-disk table fully into memory for faster subsequent access. Both.b2ddirectories and.b2zzip archives are supported transparently.
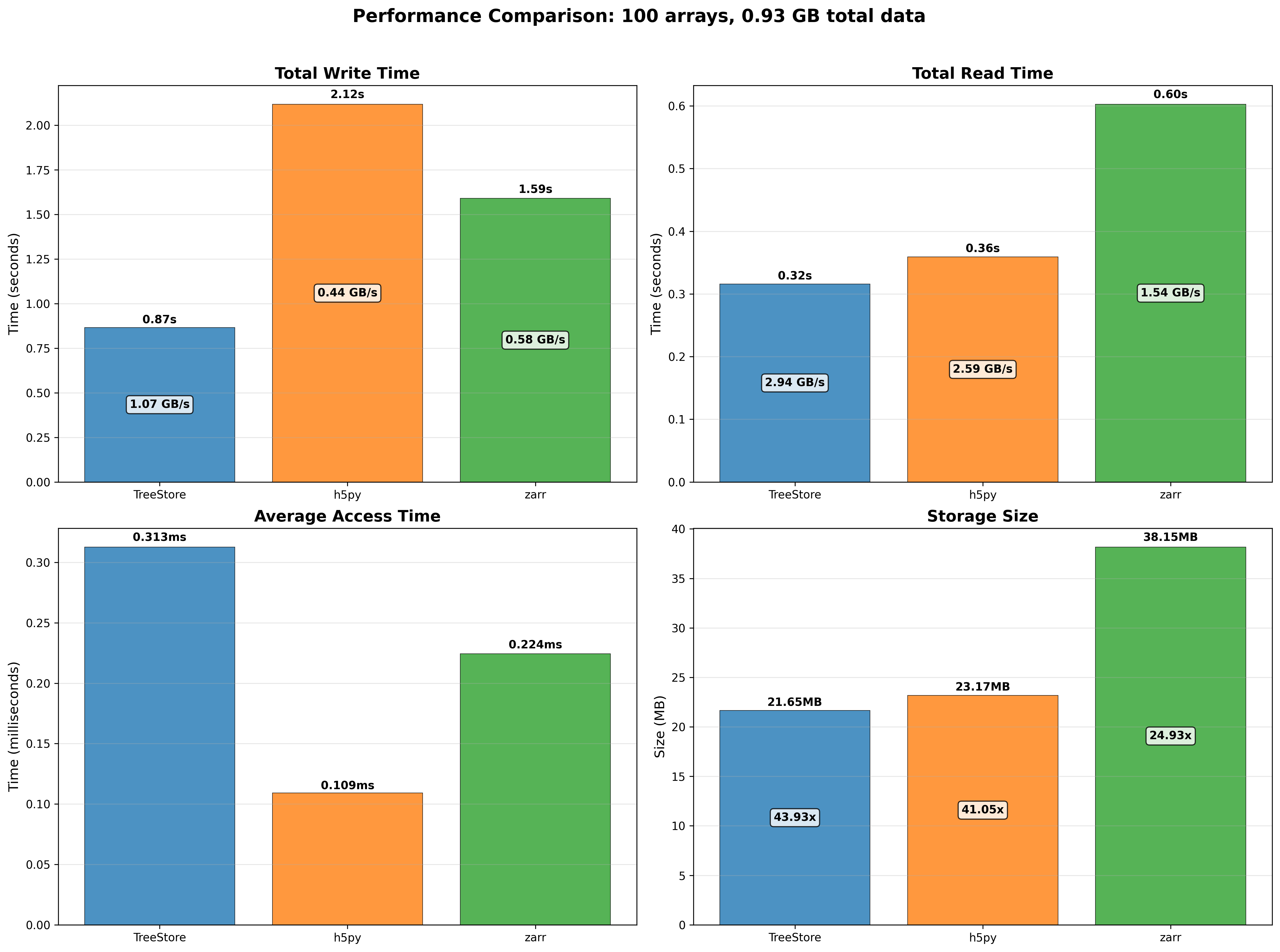
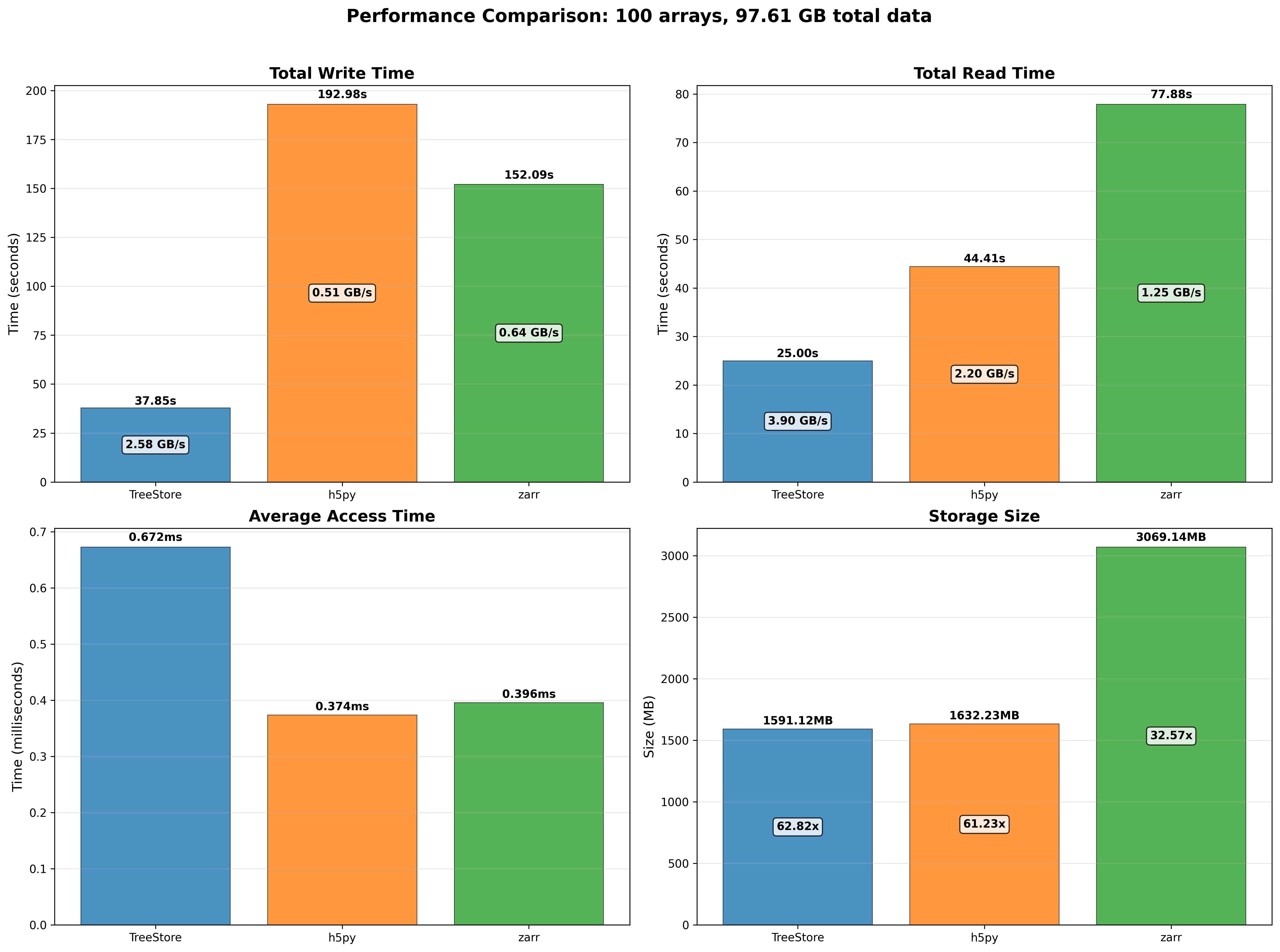
Mini benchmarks
All numbers below are from a single machine, 1 million rows, using the benchmark scripts in the repository. They are meant to give a feel for the performance characteristics, not as absolute guarantees.
Bulk loading speed
How you feed data into a CTable matters a lot. Loading 1M rows from a Python list of dicts takes around 0.66 s. Switching to a NumPy structured array brings that down to 0.03 s — a 22x speedup. Loading from an existing CTable is even faster at 28x. The takeaway is simple: if you have NumPy data, hand it directly to extend() and it will be ingested at close to raw array speed.
Filtering vs pandas
Filtering 1M rows with a range query (id between 250k and 750k, so 50% of the table) takes around 13 ms in CTable vs 31 ms in pandas — 2.4x faster. On top of that, the CTable occupies 20 MB compressed versus 31 MB for the equivalent pandas DataFrame, a 1.6x reduction in memory essentially for free thanks to Blosc2's compression pipeline.
extend() vs append() — always batch if you can
CTable has two ways to insert data: append() adds one row at a time and goes through a full Pydantic validation cycle per row; extend() takes a batch and validates it in one vectorized NumPy pass. At 100k rows the difference is 2000x in favour of ``extend()``. Even at 10k rows it is already 700x. The message is simple: if you have more than a handful of rows to insert, always batch them into a single extend() call.
where() is nearly free regardless of selectivity
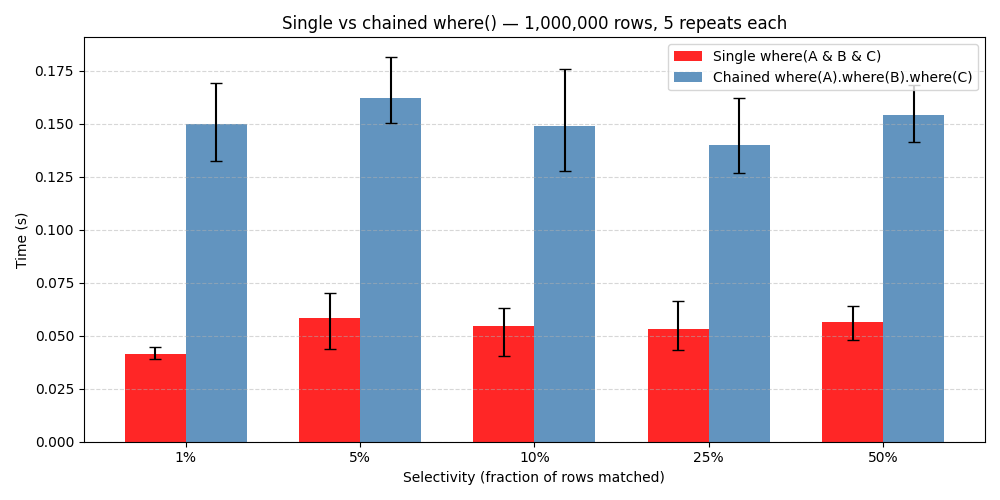
One property of the lazy mask approach is that where() costs roughly the same whether the result contains 10 rows or 999,990 rows out of 1M. In practice the time stays between 12 ms and 18 ms across all selectivity levels. You are not paying to materialise the matching rows — you are only computing a mask. The data is only read when you actually access it.
Combining filters is 4x faster than chaining them
It is tempting to filter a CTable step by step — first narrow by one condition, then filter the result by another. But each where() call creates a new view with its own mask computation. A single where() with all conditions joined by & does the same work in one pass and is 4.4x faster than five chained calls returning the same final result.
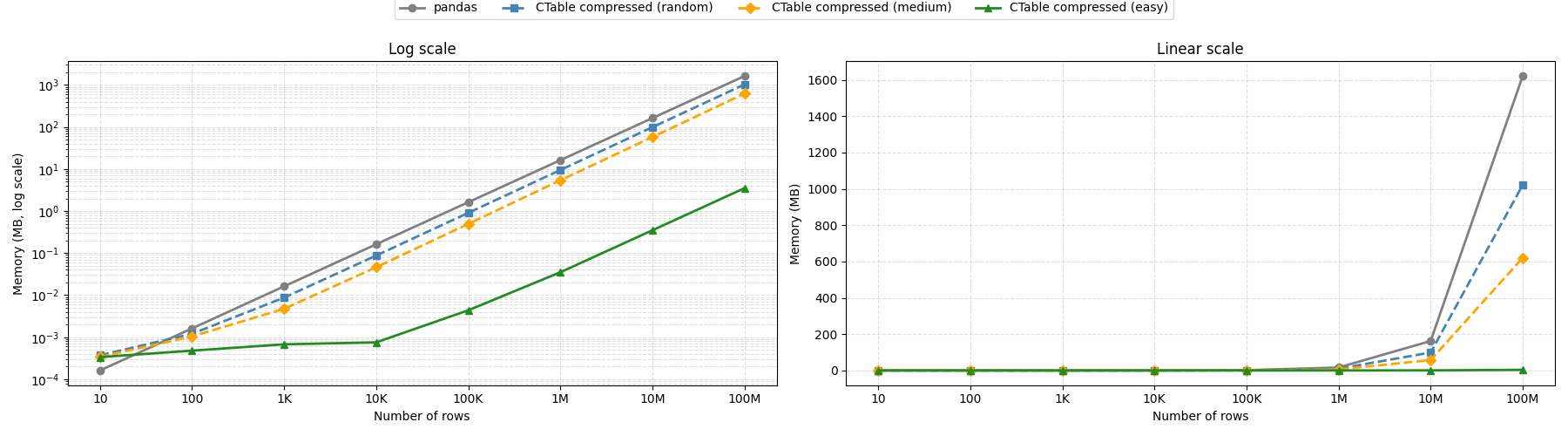
Memory footprint depends on your data
CTable compresses each column independently with Blosc2, so how much memory you save depends on how much structure your data has. With highly repetitive data — sequential integer IDs, a handful of distinct float values, constant booleans — a 100-million-row table fits in under 4 MB, versus over 1.6 GB for the equivalent pandas DataFrame. With fully random data the gain is more modest (around 1.6×), since high-entropy values leave little for the compressor to exploit. Real-world datasets typically land somewhere in between, but CTable is consistently more memory-efficient than pandas regardless of data entropy.
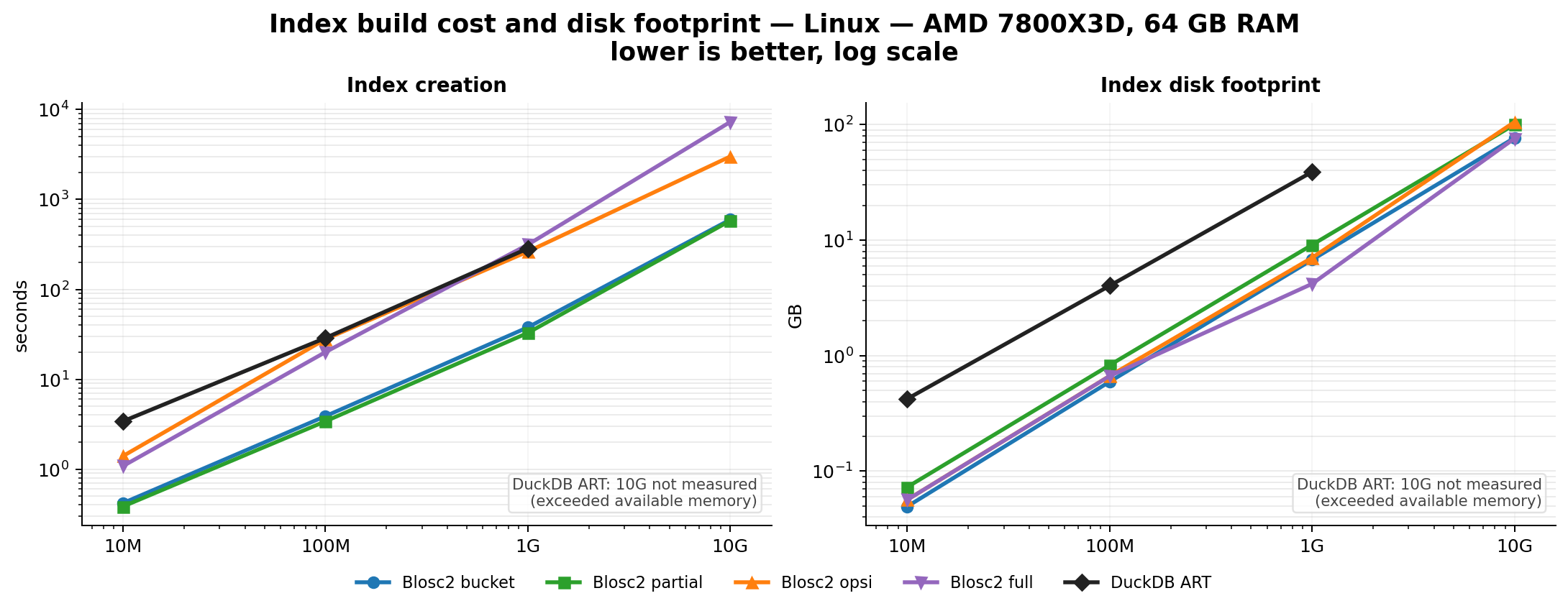
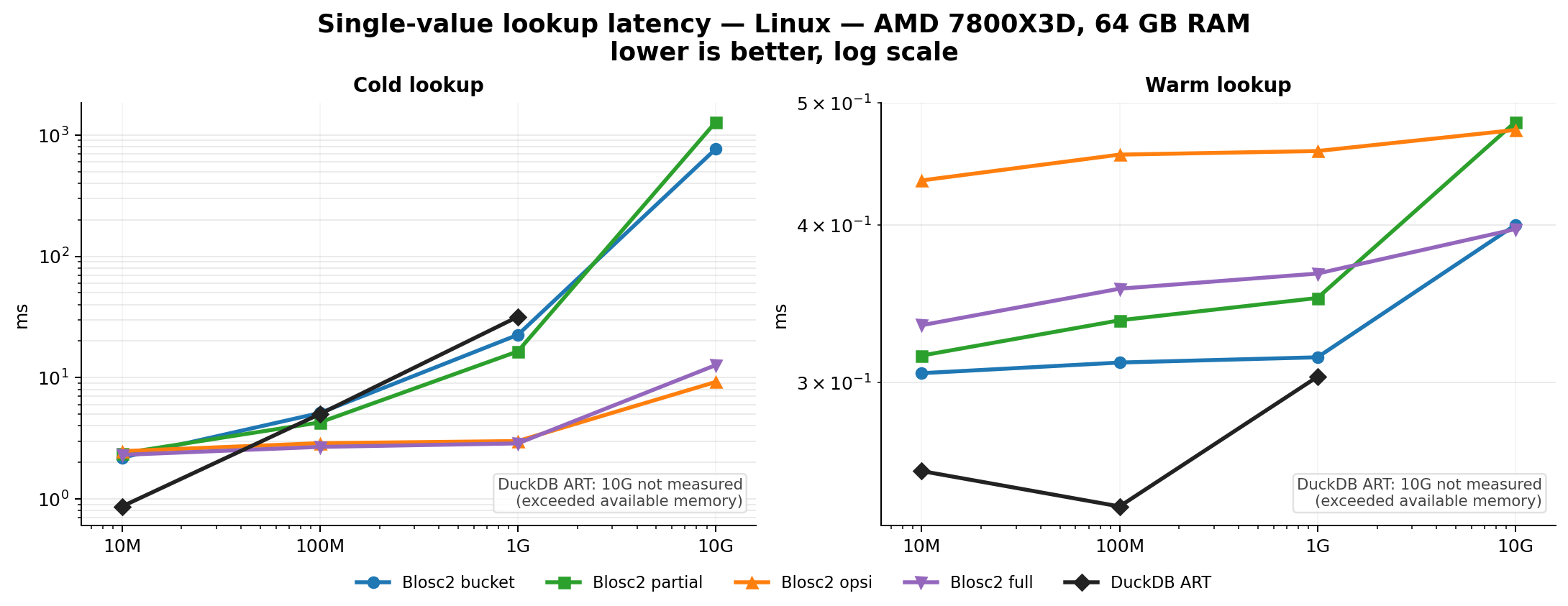
Efficient Indexing
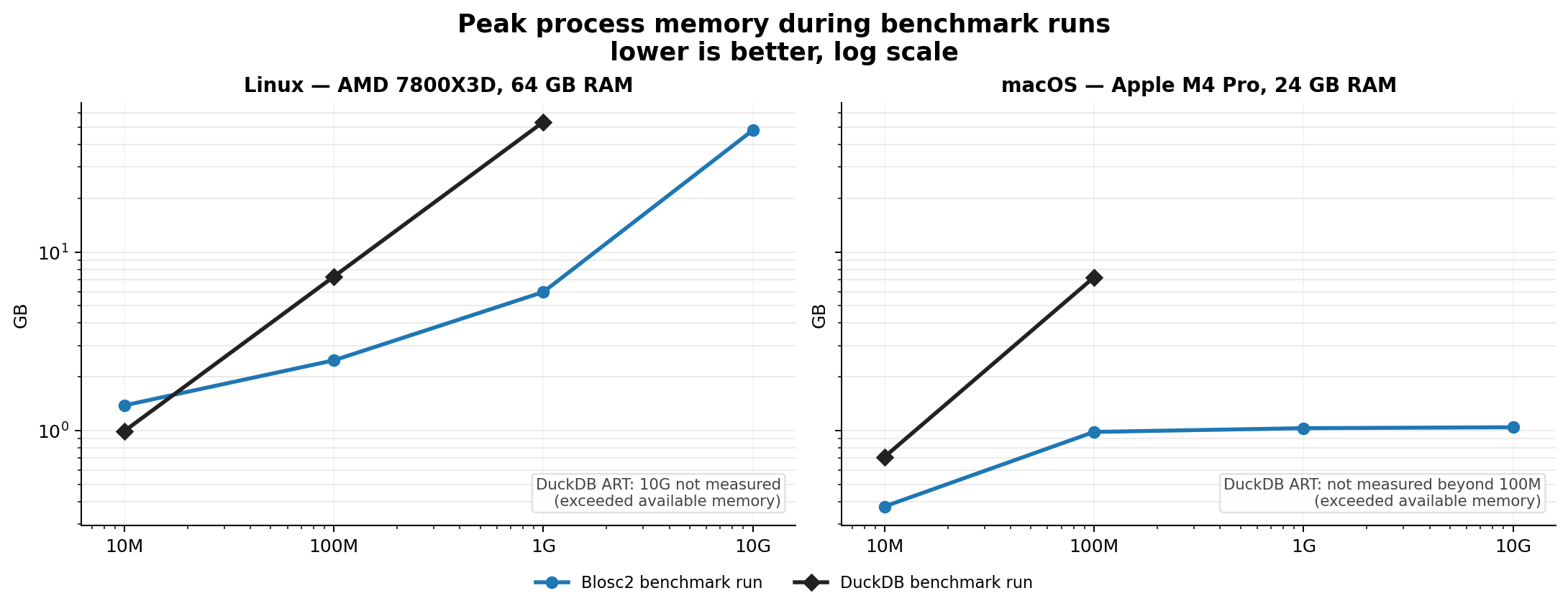
Indexes can speed up queries by orders of magnitude. The embedded indexing engine stores data in compressed form, allowing for large savings in memory and disk. For example, a FULL index on a 1 billion rows with random values takes around 5.8 GB on disk, while an equivalent DuckDB index takes around 41 GB. Typically, queries that would take seconds can be reduced to milliseconds.
Schema validation has near-zero cost at scale
Every CTable has a typed schema with optional constraints (ranges, string lengths, etc.). When inserting data with extend(), these constraints are checked via a vectorized NumPy path rather than row by row. At 1M rows with a NumPy structured array the validation overhead is essentially 1.00x —indistinguishable from skipping validation entirely. Even with Python list input it only adds 1.31x. You get correctness guarantees without paying for them at scale.
AI role
During the development of CTable, we have been using AI tools to help us in the design and implementation of the API, as well as in the documentation. Tools like Perplexity, and agents like Pi, Codex or Claude have been instrumental in helping us throughout the process, which allowed us to be much more ambitious in the features we wanted to implement. Note from Francesc: I specially liked the combination of the Pi agent and GPT 5.5 model (essentially GPT >= 5.3); that worked really well!
Of course, we already borrowed some ideas from other libraries, like Apache Arrow, Pandas, Polars, DuckDB or PyTables, but we also wanted to bring some unique features to CTable, like the ability to operate on compressed data without decompressing it, or the rich schema specs for expressing complex data types; AI has been instrumental in allowing us doing this.
Being a powerful tool, AI always need supervision and guidance to be used effectively, and we have spent lots of time bringing our cumulated decades-long experience for review code, designing tests and benchmarks, and fine-tuning the internal knobs for allowing best performance and user experience. We must say that we are very happy with the results: combining our experience with the power of AI has allowed us to create a powerful and flexible tabular data container that is well tested and documented.
More info
We have setup a couple of tutorials and a complete API reference to get you started with CTable:
Conclusions
CTable brings together compression, schema validation, and query acceleration in a self-contained Python package. It is still young, but the architecture is solid and the feature set already covers most common analytical workflows, and we hope it will be useful for many users in the Python ecosystem. We are looking forward to seeing how the community will use and contribute to Python-Blosc2 in general, an CTable in particular, and to continue improving it based on feedback and contributions from users.
Enjoy data!