Introducing Blosc2 NDim
One of the latest and more exciting additions in recently released C-Blosc2 2.7.0 is the Blosc2 NDim layer (or b2nd for short). It allows to create and read n-dimensional datasets in an extremely efficient way thanks to a completely general n-dim 2-level partitioning, allowing to slice and dice arbitrary large (and compressed!) data in a more fine-grained way.
Remember that having a second partition means that we have better flexibility to fit the different partitions at the different CPU cache levels; typically the first partition (aka chunks) should be made to fit in L3 cache, whereas the second partition (aka blocks) should rather fit in L2/L1 caches (depending on whether compression ratio or speed is desired).
This capability was formerly part of Caterva, and now we are including it in C-Blosc2 for convenience. As a consequence, the Caterva and Python-Caterva projects are now officially deprecated and all the action will happen in the C-Blosc2 / Python-Blosc2 side of the things.
Last but not least, Blosc NDim is gaining support for a full-fledged data type system like NumPy. Keep reading.
Going multidimensional in the first and the second partition
Blosc (both Blosc1 and Blosc2) has always had a two-level partition schema to leverage the different cache levels in modern CPUs and make compression happen as quickly as possible. This allows, among other things, to create and query tables with 100 trillion of rows when properly cooperating with existing libraries like HDF5.
With Blosc2 NDim we are taking this feature a step further and both partitions, known as chunks and blocks, are gaining multidimensional capabilities meaning that one can split some dataset (super-chunk in Blosc2 parlance) in such a n-dim cubes and sub-cubes:
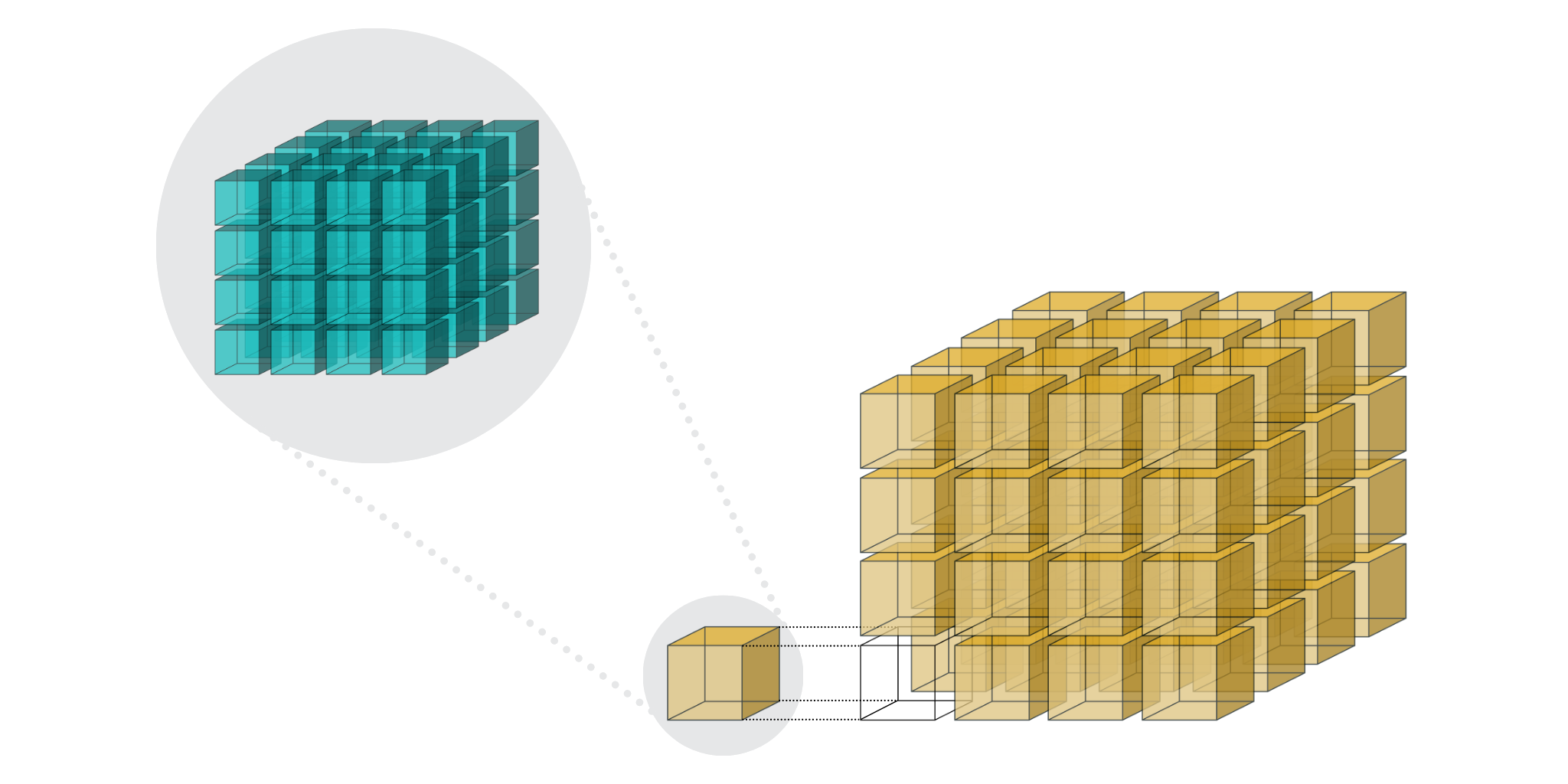
With these more fine-grained cubes (aka partitions), it is possible to retrieve arbitrary n-dim slices more rapidly because you don't have to decompress all the data that is necessary for the more coarse-grained partitions typical in other libraries.
For example, for a 4-d array with a shape of (50, 100, 300, 250) with float64 items, we can choose a chunk with shape (10, 25, 50, 50) and a block with shape (3, 5, 10, 20) which makes for about 5 MB and 23 KB respectively. This way, a chunk fits comfortably on a L3 cache in most of modern CPUs, and a block in a L1 cache (we are tuning for speed here). With that configuration, the NDArray object in the Python-Blosc2 package can slice the array as fast as it is shown below:
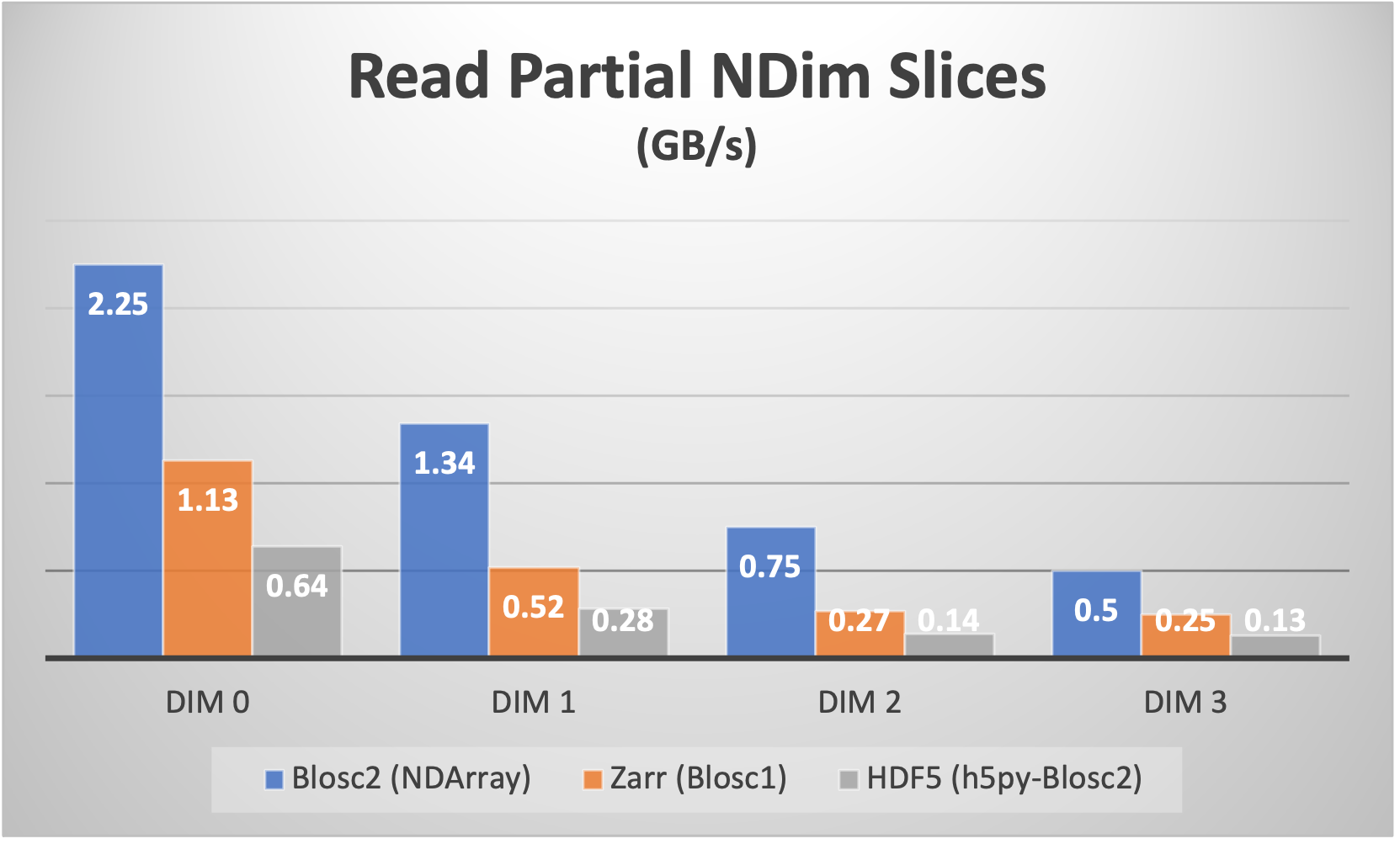
Of course, the double partition comes with some overhead during the creation of the partitions: more data moves and computations are required in order to place the data in the correct positions. However, we have done our best in order to minimize the data movement as much as possible. Below we can see how the speed of creation (write) of an array from anew is still quite competitive:
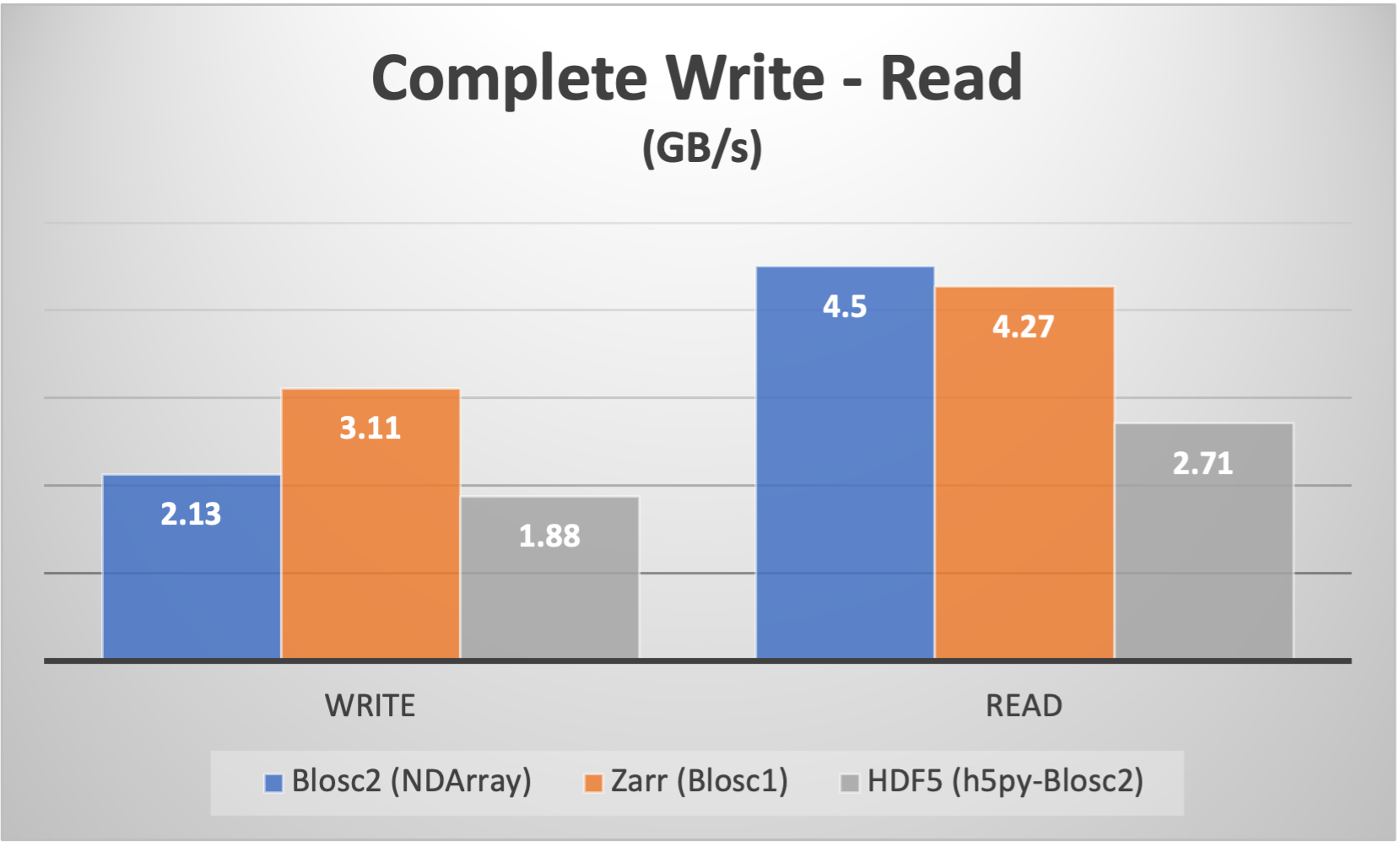
On the other hand, we can also see that, when reading the complete array, the double partitioning overhead is not really a big issue, and actually, it benefits Blosc2 NDArray somewhat.
All the plots above have been generated using the compare_getslice.py script, where we have been using the Zstd codec with compression level 1 (the fastest inside Blosc2) + the Shuffle filter for all the packages. The box used was an Intel 13900K CPU with 32 GB of RAM and using an up-to-date Clear Linux distro.
Data types are in!
Another important thing that we are adding to Blosc2 NDim is the support for data types. This was not previously supported in either C-Blosc2 or Caterva, where only a typesize was available to characterize the type. Now, the data type becomes a first class citizen for the b2nd metalayer. Metalayers in Blosc2 are stored in msgpack format, so it is pretty easy to introspect into them by using external msgpack tools. For example, the b2nd file created in the section above contains this meta info:
$ dd bs=1 skip=112 count=1000 < compare_getslice.b2nd | msgpack2json -b <snip> [0,4,[50,100,300,250],[10,25,50,50],[3,5,10,20],0,"<f8"]
Here we can see the version of the metalayer (0), the number of dimensions of the array (4), followed by the shape, chunk shape and block shape. Then it comes the version of the dtype representation (it support up to 127; the default is 0, meaning NumPy). Finally, we can spot the "<f8" string, so a little-endian double precision data type. Note that the all data types in NumPy are supported by the Python wrapper of Blosc2; that means that with the NDArray object you can store e.g. datetimes (including units), or arbitrarily nested heterogeneous types, which allows to create multidimensional tables.
Conclusion
We have seen how, when sensibly chosen, the double partition provides a formidable boost in retrieving arbitrary slices in potentially large multidimensional arrays. In addition, the new support for arbitrary data types represents a powerful addition as well. Combine that with the excellent compression capabilities of Blosc2, and you will get a first class data container for many types of (numerical, but also textual) data.
Finally, we will be releasing the new NDArray object in the forthcoming release of Python-Blosc2 very soon. This will enable full access to these shiny new features of Blosc2 from the convenience of Python. Stay tuned!
If you regularly store and process large datasets and need advice to partition your data, or choosing the best combination of codec, filters, chunk and block sizes, or many other aspects of compression, do not hesitate to contact the Blosc team at contact (at) blosc.org. We have more than 30 years of cumulative experience in data handling systems like HDF5, Blosc and efficient I/O in general; but most importantly, we have the ability to integrate these innovative technologies quickly into your products, enabling a faster access to these innovations.
Update (2023-02-24)
We have released Python-Blosc2 2.1.1 with the new NDArray object: https://github.com/Blosc/python-blosc2/releases/
We have also prepared a ~2 min explanatory video on why slicing in a pineapple-style (aka double partition) is useful:

Enjoy the meal!
Comments
Comments powered by Disqus